Note
Go to the end to download the full example code
Vector Data#
Some datasets have multiple vector components measured for each location, like the East and West components of wind speed or GPS velocities. For example, let’s look at our sample GPS velocity data from the U.S. West coast.
import cartopy.crs as ccrs
import matplotlib.pyplot as plt
import pyproj
import verde as vd
data = vd.datasets.fetch_california_gps()
# We'll need to project the geographic coordinates to work with our Cartesian
# classes/functions
projection = pyproj.Proj(proj="merc", lat_ts=data.latitude.mean())
proj_coords = projection(data.longitude.values, data.latitude.values)
# This will be our desired grid spacing in degrees
spacing = 12 / 60
plt.figure(figsize=(6, 8))
ax = plt.axes(projection=ccrs.Mercator())
crs = ccrs.PlateCarree()
tmp = ax.quiver(
data.longitude.values,
data.latitude.values,
data.velocity_east.values,
data.velocity_north.values,
scale=0.3,
transform=crs,
width=0.002,
)
ax.quiverkey(tmp, 0.2, 0.15, 0.05, label="0.05 m/yr", coordinates="figure")
ax.set_title("GPS horizontal velocities")
vd.datasets.setup_california_gps_map(ax)
plt.show()
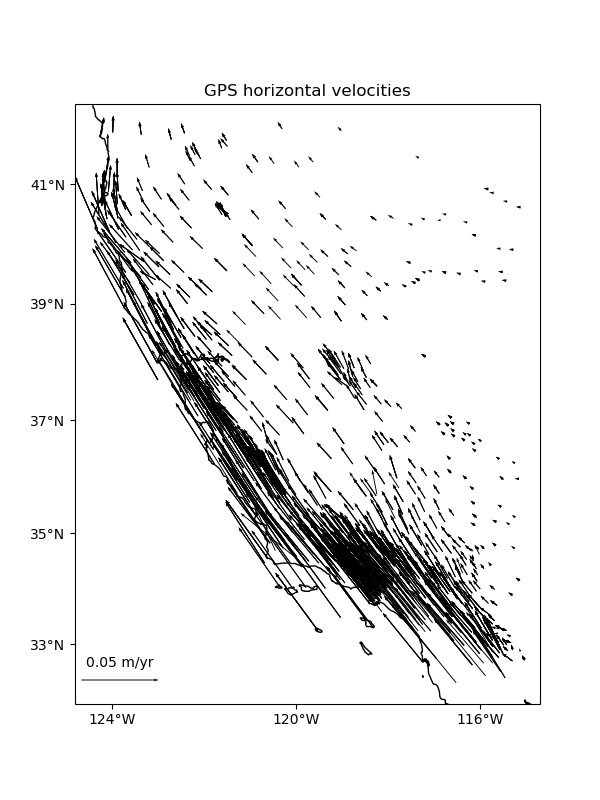
Verde classes and functions are equipped to deal with vector data natively or
through the use of the verde.Vector class. Function and classes that
can take vector data as input will accept tuples as the data and
weights arguments. Each element of the tuple must be an array with the
data values for a component of the vector data. As with coordinates,
the order of components must be (east_component, north_component,
up_component).
Blocked reductions#
Operations with verde.BlockReduce and verde.BlockMean can
handle multi-component data automatically. The reduction operation is applied
to each data component separately. The blocked data and weights will be
returned in tuples as well following the same ordering as the inputs. This
will work for an arbitrary number of components.
# Use a blocked mean with uncertainty type weights
reducer = vd.BlockMean(spacing=spacing * 111e3, uncertainty=True)
block_coords, block_data, block_weights = reducer.filter(
coordinates=proj_coords,
data=(data.velocity_east, data.velocity_north),
weights=(1 / data.std_east**2, 1 / data.std_north**2),
)
print(len(block_data), len(block_weights))
2 2
We can convert the blocked coordinates back to longitude and latitude to plot with Cartopy.
block_lon, block_lat = projection(*block_coords, inverse=True)
plt.figure(figsize=(6, 8))
ax = plt.axes(projection=ccrs.Mercator())
crs = ccrs.PlateCarree()
tmp = ax.quiver(
block_lon,
block_lat,
block_data[0],
block_data[1],
scale=0.3,
transform=crs,
width=0.002,
)
ax.quiverkey(tmp, 0.2, 0.15, 0.05, label="0.05 m/yr", coordinates="figure")
ax.set_title("Block mean velocities")
vd.datasets.setup_california_gps_map(ax)
plt.show()
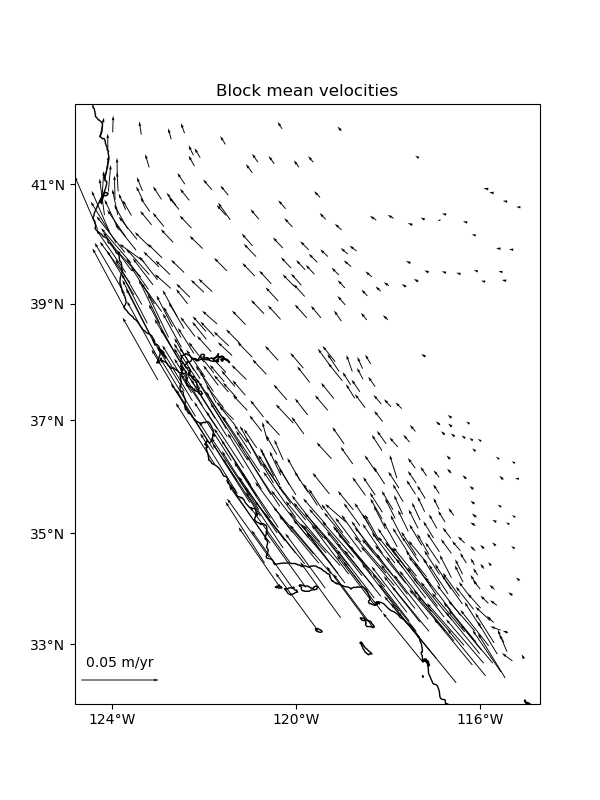
Trends#
Trends can’t handle vector data automatically, so you can’t pass
data=(data.velocity_east, data.velocity_north) to
verde.Trend.fit. To get around that, you can use the
verde.Vector class to create multi-component estimators and gridders
from single component ones.
Vector takes an estimator/gridder for each data component and
implements the gridder interface for vector data,
fitting each estimator/gridder given to a different component of the data.
For example, to fit a trend to our GPS velocities, we need to make a 2-component vector trend:
Vector(components=[Trend(degree=4), Trend(degree=1)])
We can use the trend as if it were a regular verde.Trend but
passing in 2-component data to fit. This will fit each data component to a
different verde.Trend.
trend.fit(
coordinates=proj_coords,
data=(data.velocity_east, data.velocity_north),
weights=(1 / data.std_east**2, 1 / data.std_north**2),
)
Each estimator can be accessed through the components attribute:
print(trend.components)
print("East trend coefficients:", trend.components[0].coef_)
print("North trend coefficients:", trend.components[1].coef_)
[Trend(degree=4), Trend(degree=1)]
East trend coefficients: [-3.31743842e+03 -1.61368024e-03 -1.08568612e-03 -2.77735425e-10
-2.99424188e-10 7.38718818e-12 -2.03916793e-17 -2.68710016e-17
3.36222190e-18 2.00921241e-18 -5.43959103e-25 -7.79030546e-25
2.71546631e-25 2.44086928e-25 5.03611061e-26]
North trend coefficients: [-3.35415183e-01 -4.50275691e-08 -3.75590521e-08]
When we call verde.Vector.predict or verde.Vector.grid, we’ll
get back predictions for two components instead of just one. Each prediction
comes from a different verde.Trend.
pred_east, pred_north = trend.predict(proj_coords)
# Make histograms of the residuals
plt.figure(figsize=(6, 5))
ax = plt.axes()
ax.set_title("Trend residuals")
ax.hist(data.velocity_north - pred_north, bins="auto", label="North", alpha=0.7)
ax.hist(data.velocity_east - pred_east, bins="auto", label="East", alpha=0.7)
ax.legend()
ax.set_xlabel("Velocity (m/yr)")
plt.show()
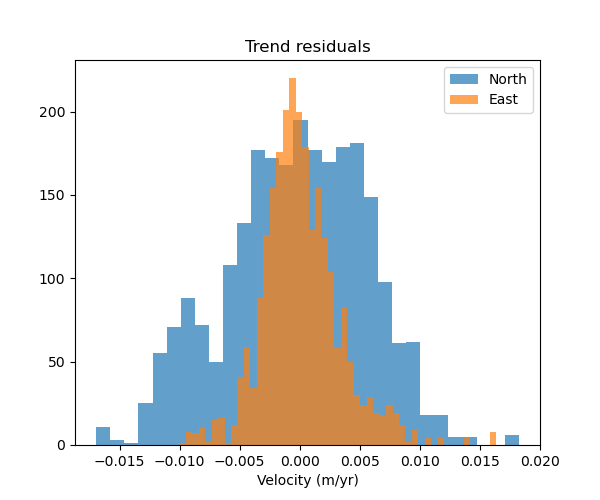
As expected, the residuals are higher for the North component because of the lower degree polynomial.
Let’s make geographic grids of these trends.
region = vd.get_region((data.longitude, data.latitude))
grid = trend.grid(
region=region,
spacing=spacing,
projection=projection,
dims=["latitude", "longitude"],
)
print(grid)
<xarray.Dataset> Size: 38kB
Dimensions: (latitude: 49, longitude: 47)
Coordinates:
* latitude (latitude) float64 392B 32.29 32.49 32.69 ... 41.7 41.9
* longitude (longitude) float64 376B 235.7 235.9 236.1 ... 244.8 245.0
Data variables:
east_component (latitude, longitude) float64 18kB -0.1983 ... 0.01365
north_component (latitude, longitude) float64 18kB 0.0541 ... -0.02427
Attributes:
metadata: Generated by Vector(components=[Trend(degree=4), Trend(degree=...
By default, the names of the data components in the xarray.Dataset
are east_component and north_component. This can be customized using
the data_names argument.
Now we can map the trends.
fig, axes = plt.subplots(
1, 2, figsize=(9, 7), subplot_kw=dict(projection=ccrs.Mercator())
)
crs = ccrs.PlateCarree()
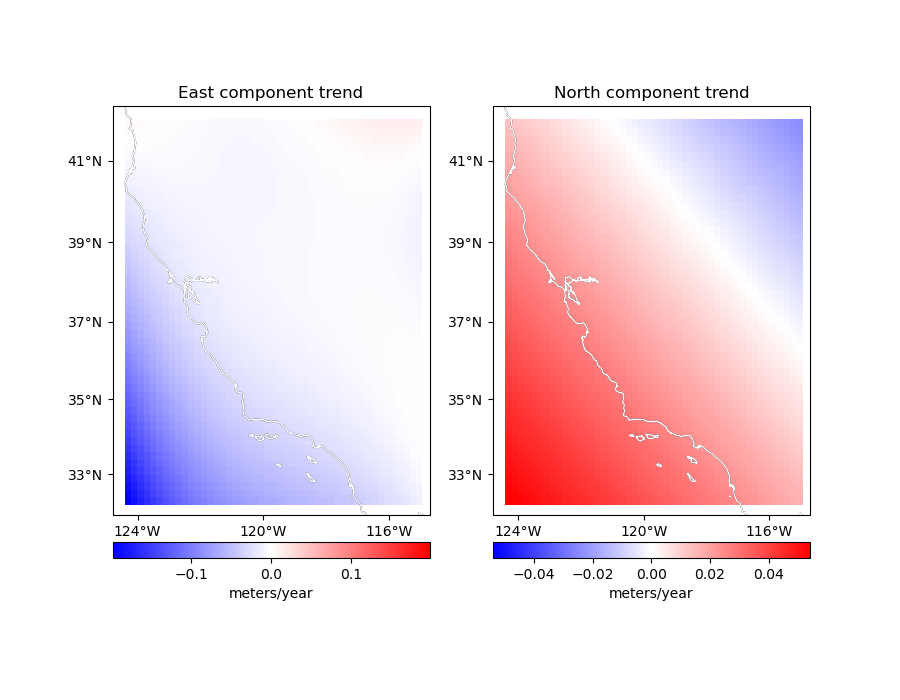
titles = ["East component trend", "North component trend"]
components = [grid.east_component, grid.north_component]
for ax, component, title in zip(axes, components, titles):
ax.set_title(title)
maxabs = vd.maxabs(component)
tmp = component.plot.pcolormesh(
ax=ax,
vmin=-maxabs,
vmax=maxabs,
cmap="bwr",
transform=crs,
add_colorbar=False,
add_labels=False,
)
cb = plt.colorbar(tmp, ax=ax, orientation="horizontal", pad=0.05)
cb.set_label("meters/year")
vd.datasets.setup_california_gps_map(ax, land=None, ocean=None)
ax.coastlines(color="white")
plt.show()
/home/runner/work/verde/verde/doc/tutorials_src/vectors.py:197: UserWarning: All kwargs are being ignored. They are accepted to guarantee backward compatibility.
vd.datasets.setup_california_gps_map(ax, land=None, ocean=None)
/home/runner/work/verde/verde/doc/tutorials_src/vectors.py:197: UserWarning: All kwargs are being ignored. They are accepted to guarantee backward compatibility.
vd.datasets.setup_california_gps_map(ax, land=None, ocean=None)
Gridding#
You can use verde.Vector to create multi-component gridders out of
verde.Spline the same way as we did for trends. In this case, each
component is treated separately.
We can start by splitting the data into training and testing sets (see
Model Selection). Notice that verde.train_test_split work for
multicomponent data automatically.
train, test = vd.train_test_split(
coordinates=proj_coords,
data=(data.velocity_east, data.velocity_north),
weights=(1 / data.std_east**2, 1 / data.std_north**2),
random_state=1,
)
Now we can make a 2-component spline. Since verde.Vector implements
fit, predict, and filter, we can use it in a verde.Chain
to build a pipeline.
We need to use a bit of damping so that the weights can be taken into account. Splines without damping provide a perfect fit to the data and ignore the weights as a consequence.
Chain(steps=[('mean', BlockMean(spacing=22200.0, uncertainty=True)),
('trend', Vector(components=[Trend(degree=1), Trend(degree=1)])),
('spline',
Vector(components=[Spline(damping=1e-10, mindist=0),
Spline(damping=1e-10, mindist=0)]))])
Warning
Never generate the component gridders with [vd.Spline()]*2. This will
result in each component being a represented by the same Spline
object, causing problems when trying to fit it to different components.
Fitting the spline and gridding is exactly the same as what we’ve done before.
/home/runner/work/verde/verde/doc/tutorials_src/vectors.py:252: FutureWarning: The default scoring will change from R² to negative root mean squared error (RMSE) in Verde 2.0.0. This may change model selection results slightly.
print(chain.score(*test))
0.9926765596607829
<xarray.Dataset> Size: 38kB
Dimensions: (latitude: 49, longitude: 47)
Coordinates:
* latitude (latitude) float64 392B 32.29 32.49 32.69 ... 41.7 41.9
* longitude (longitude) float64 376B 235.7 235.9 236.1 ... 244.8 245.0
Data variables:
east_component (latitude, longitude) float64 18kB -0.0466 ... -0.0006105
north_component (latitude, longitude) float64 18kB 0.08795 ... -0.0005965
Attributes:
metadata: Generated by Chain(steps=[('mean', BlockMean(spacing=22200.0, ...
Mask out the points too far from data and plot the gridded vectors.
grid = vd.distance_mask(
(data.longitude, data.latitude),
maxdist=spacing * 2 * 111e3,
grid=grid,
projection=projection,
)
plt.figure(figsize=(6, 8))
ax = plt.axes(projection=ccrs.Mercator())
tmp = ax.quiver(
grid.longitude.values,
grid.latitude.values,
grid.east_component.values,
grid.north_component.values,
scale=0.3,
transform=crs,
width=0.002,
)
ax.quiverkey(tmp, 0.2, 0.15, 0.05, label="0.05 m/yr", coordinates="figure")
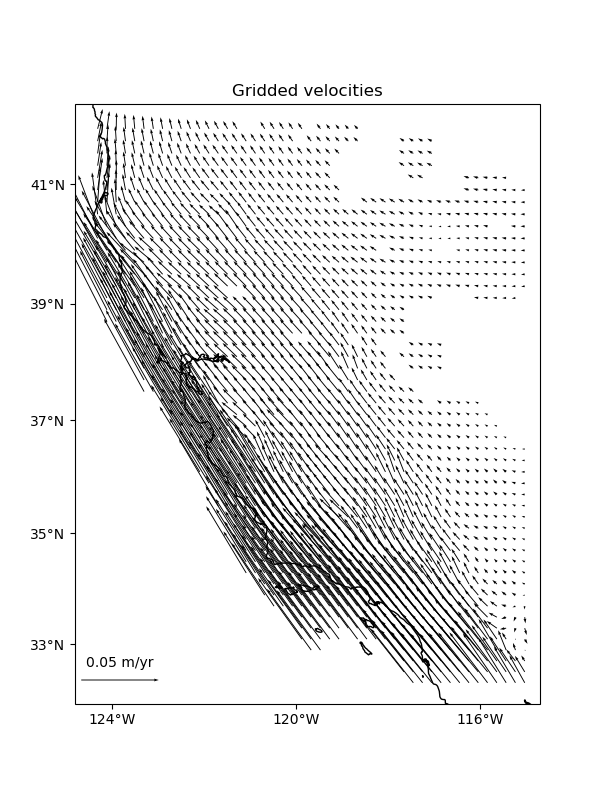
ax.set_title("Gridded velocities")
vd.datasets.setup_california_gps_map(ax)
plt.show()
GPS/GNSS data#
For some types of vector data, like GPS/GNSS displacements, the vector
components are coupled through elasticity. In these cases, elastic Green’s
functions can be used to achieve better interpolation results. The
verde.VectorSpline2D implements these Green’s functions.
Total running time of the script: (0 minutes 1.005 seconds)